Humans and domesticated animals
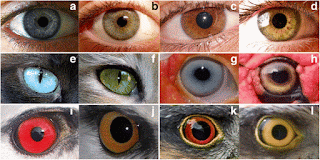
tend to show variation in eye color unlike in wild animals whose eye colors is usually
fixed. It is however not known when this variation in eye color emerged in the
evolution of the genus Homo and domesticated animals. From previous research,
we know however that the emergence and fixation of variants in both coat, plumage
and eye coloration started during the early stages of domestication in the
Neolithic. Eye coloration in humans is known to be continuous having numerous
shades ranging from different shades of light blue to dark brown. The very rare
cases of eye coloration in wild animals is associated with maturation with age
as seen in birds and some instances of sexual dimorphism (certain duck
species). Who knows perhaps in humans it is a case of sexual selection with
blue and green eyed individuals preferred more as mates than brown eyed
individuals.
News Article: here
Research Article: here
Joanne








/cdn.vox-cdn.com/uploads/chorus_asset/file/9250027/image1.png)













